Motifs and Domains:
Two important distinguishing features to look for when studying proteins are motifs and domains. A motif, sometimes referred to as a sequence feature, is the sequence of amino acids (the building blocks of a protein) that are responsible for a certain type of activity of a protein. For example, an important motif for an enzyme would be the amino acids in the active site responsible for that enzyme’s catalytic activity. A protein domain is a distinct functional unit of a protein accounting for a specific function of that protein. Domains can be used to classify proteins into family hierarchs allowing for the assortment of proteins into groups. This is important because it allows us to predict the function of a novel protein based on the group it is thought to be a part of [1]. This also allows for us to determine how similar two putative homologs are in function based on presence of absence of certain domains that confer the function of the protein of interest.
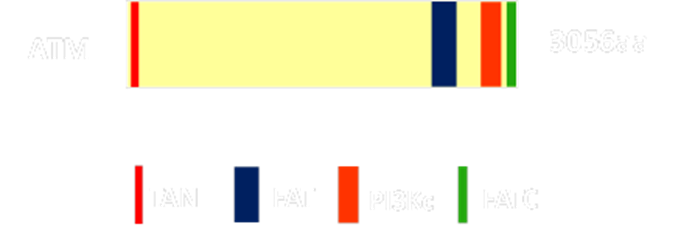
The figure bellow shows the known motifs/domains of the Ataxia Telangiectasia Mutated (ATM) protein. These domains were found by subjecting the human ATM protein sequence to PFAM and SMART databases and the resulting data was used to construct the figure using power point.
The figure bellow shows the known motifs/domains of the Ataxia Telangiectasia Mutated (ATM) protein. These domains were found by subjecting the human ATM protein sequence to PFAM and SMART databases and the resulting data was used to construct the figure using power point.
Figure 1: The Human ATM protein and its known domains.
PI3Kc: This domain has amino acids that are similar to those found in catalytic subunit of phosphatidylinositol 3-kinases, better known as PI 3-kinases. A kinase is an enzyme that transfers a phosphate group onto a different protein, which can, in turn, activate or inhibit that protein. PI 3-kinases are given this name because they transfer a phosphate group to the phospholipid phosphatidylinositol, abbreviated PI [2]. Proteins that carry domains similar to this one, more commonly termed PI 3-kinase-related regions, are usually involved in cell cycle regulation, responses to DNA damage and maintenance of genome stability [3]. Considering this information it is likely that the majority of ATM’s function is attributed to this domain.
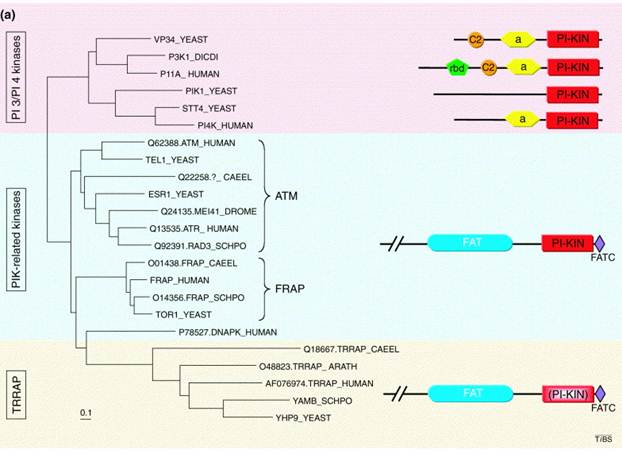
FAT and FATC: The FAT domain got its name due to the fact that it is only found in the FRAP ATM and TRRAP protein subfamilies (Figure 2). This domain always is seen with the FATC domain which is why it is speculated that they might interact together be forming a pocket that allows the PI3Kc domain to work properly. Though there is more research that needs to be done, it is hypothesized that the FAT domain serves as a structural scaffold or as a protein-binding domain or maybe even both [4].
TAN: This motif, located at the N-terminus end of the protein, has been found in ATM orthologs of vertebrates, flies, and yeast. Though plants have been shown to have the TAN motif too, it is unclear whether or not it is present in Arabidopsis thaliana. The function of this motif was studied using the yeast homolog Tel1 and it was found to be important in telomere length maintenance as well as playing a key role in the recruitment of Tel1 near double stranded breaks. The fact that this motif shows a high level of conservation it is likely that it serves in similar functions in other organisms as well [5].
FAT and FATC: The FAT domain got its name due to the fact that it is only found in the FRAP ATM and TRRAP protein subfamilies (Figure 2). This domain always is seen with the FATC domain which is why it is speculated that they might interact together be forming a pocket that allows the PI3Kc domain to work properly. Though there is more research that needs to be done, it is hypothesized that the FAT domain serves as a structural scaffold or as a protein-binding domain or maybe even both [4].
TAN: This motif, located at the N-terminus end of the protein, has been found in ATM orthologs of vertebrates, flies, and yeast. Though plants have been shown to have the TAN motif too, it is unclear whether or not it is present in Arabidopsis thaliana. The function of this motif was studied using the yeast homolog Tel1 and it was found to be important in telomere length maintenance as well as playing a key role in the recruitment of Tel1 near double stranded breaks. The fact that this motif shows a high level of conservation it is likely that it serves in similar functions in other organisms as well [5].
Figure 2: This is a tree showing how only ATM, FRAP and TRRAP proteins have a FAT domain [3].
discussion:
By using PFAM and SMART it was determined that all the homologs of ATM found in common model organisms contained the PI3kc and FATC domains. Frogs (Xenopus Laevis), mice, and zebra fish harbor all the domains of the human ATM protein, while C.elegans and Drosophila only possessed the PI3Kc and FATC domains. Yeast’s homolog of ATM had the PI3Kc and FATC domains as well as the TAN motif. The fact that ATM has been conserved across evolutionary time is an indication of its importance within the cell. Furthermore, given that ATM homologs contain the same core domain (PI3Kc and FATC) means that there are a plethora of research options available when studying ATM. Using organisms such as mice and zebra fish can be useful when trying to better understand ATM’s function in human diseases, while doing experiments with lower organisms, such as C.elegans and Drosophila, can help delineate how ATM has evolved.
For a figure depicting the homologs of ATM in various organisms, click on the file below. The information for this figure was obtained by using the PFAM and SMART domains and then was created using power point.
For a figure depicting the homologs of ATM in various organisms, click on the file below. The information for this figure was obtained by using the PFAM and SMART domains and then was created using power point.
| atm_homologs.pdf | |
| File Size: | 94 kb |
| File Type: | |
References:
1. http://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification (This site gives a brief tutorial on classifying domains of proteins)
2. Keith, C. T., & Schreiber, S. L. (1995). PIK-Related Kinases: DNA Repair, Recombination, and Cell Cycle Checkpoints. Science, 270(5233), 50-51. doi:10.2307/2888208
3. Rotman, G., & Shiloh, Y. (1998). ATM: From gene to function. Human molecular genetics, 7(10), 1555-1563. doi:10.1093/hmg/7.10.1555
4. Bosotti, R., Isacchi, A., & Sonnhammer, E. L. (2000). FAT: a novel domain in PIK-related kinases. Trends in Biochemical Sciences, 25(5), 225-227. Retrieved from http://www.sciencedirect.com.ezproxy.library.wisc.edu/science/article/pii/S0968000400015632#
5. Seidel, J. J., Anderson, C. M., & Blackburn, E. H. (2008). A Novel Tel1/ATM N-Terminal Motif, TAN, Is Essential for Telomere Length Maintenance and a DNA Damage Response. Molecular and Cellular Biology, 28(18), 5736-5746. doi:10.1128/MCB.00326-08
2. Keith, C. T., & Schreiber, S. L. (1995). PIK-Related Kinases: DNA Repair, Recombination, and Cell Cycle Checkpoints. Science, 270(5233), 50-51. doi:10.2307/2888208
3. Rotman, G., & Shiloh, Y. (1998). ATM: From gene to function. Human molecular genetics, 7(10), 1555-1563. doi:10.1093/hmg/7.10.1555
4. Bosotti, R., Isacchi, A., & Sonnhammer, E. L. (2000). FAT: a novel domain in PIK-related kinases. Trends in Biochemical Sciences, 25(5), 225-227. Retrieved from http://www.sciencedirect.com.ezproxy.library.wisc.edu/science/article/pii/S0968000400015632#
5. Seidel, J. J., Anderson, C. M., & Blackburn, E. H. (2008). A Novel Tel1/ATM N-Terminal Motif, TAN, Is Essential for Telomere Length Maintenance and a DNA Damage Response. Molecular and Cellular Biology, 28(18), 5736-5746. doi:10.1128/MCB.00326-08
This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison