Homology:
Homology is the study of similarity between different organisms due to a common ancestor. Therefore, if two proteins are said to be homologous then that means they share a common ancestor [1]. There are a couple ways in which homologs can be found. One way is by using a site known as homologene in which you enter your protein name and it gives you species in which homologs are known [2]. Another way is by using BLAST, which finds homologs by taking the sequence of interest and running it through a program that will align it to other known sequences [3]. When comparing homologs it is important to take into account the identity and similarity shared between the two aligned sequences. Identity refers to the amount of identical matches found when aligning your protein of interest with the sequence from the same protein in another organism. Similarity differs from identity in that the amino acid in one position doesn’t have to be identical to the amino acid in the corresponding position on the other protein. This is because different amino acids can have similar chemical properties such as being hydrophobic or hydrophilic (two important properties when considering protein folding). Therefore if two amino acids are similar, their difference will not cause great change to the protein.
Homology is important because it allows us to look to other organisms when trying to do experiments on a protein of interest in humans. By characterizing how this protein function in other organisms we can better understand how it functions in humans.
Homology is important because it allows us to look to other organisms when trying to do experiments on a protein of interest in humans. By characterizing how this protein function in other organisms we can better understand how it functions in humans.
ATM Homologs:
The homologs for ATM were found by using the site homologene as well as doing a gene search on NCBI [2,4]. Once the homologs were found, BLAST was used to determine the identity and similarity between the various homologs. The information regarding chimps, mouse, rat, cattle, dog, cat, African-clawed frog, zebra fish, C.elegans, yeast, Arabidopsis, and fruit fly are located bellow. These were also the organisms termed "model organisms" on the protein phylogeny page. Information regarding the rest of the homologs shown can be downloaded by clicking on the graph. This information is arranged in alphabetical order according to the common species name.
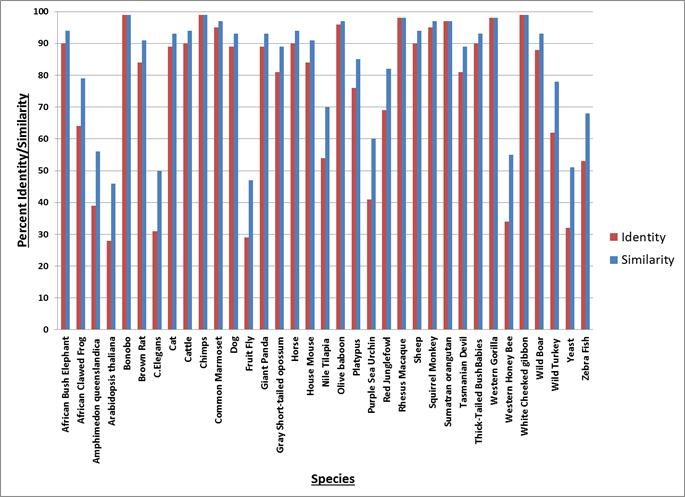
Figure 1: This graph shows how much identity (red) and similarity (blue) each organism has with the human ATM protein sequence. The species are arranged in alphabetical order.
Discussion:
Overall, it seems that the ATM gene serves an important role within all organisms as homologs of the gene were found in a wide variety or organisms. Findings of homologs in species such as Arabidopsis and yeast further support this claim as these species split off from humans and the rest of the animal kingdom long ago. Seeing as ATM is an important gene in DNA repair, this result was anticipated because all organisms need to be able repair DNA damage as it arises. The general trend shown here is that the more closely related the organisms are to humans, higher amount of identity/similarity they have with the human ATM protein. This is evident by the fact that the monkey species show the highest amount of identity/similarity with humans.
Homo Sapiens
|
Danio Rerio (Zebra Fish)
|
References:
1. Reeck, G. R., de Haen, C, Teller, D. C., Doolittle, R. F., Fitch, W. M., Dickerson, R. E., Chambon, P., McLachlan, A. D., Margoliash, E., Jukes, T. H., Zuckerandl E. (1987) “Homology” in proteins and nucleic acids: a terminology muddle and a way out of it. Cell, 1987(50), 667. doi: 10.1016/0092-8674(87)90322-9
2. http://www.ncbi.nlm.nih.gov/homologene (Homologene)
3. Dereeper A., Audic S., Claverie J., Blanc G. BLAST-EXPLORER helps you building datasets for phylogenetic analysis. BMC Evol Biol. 2010 Jan 12;10:8. PMID:20067610
4. http://www.ncbi.nlm.nih.gov/gene (NCBI gene search)
2. http://www.ncbi.nlm.nih.gov/homologene (Homologene)
3. Dereeper A., Audic S., Claverie J., Blanc G. BLAST-EXPLORER helps you building datasets for phylogenetic analysis. BMC Evol Biol. 2010 Jan 12;10:8. PMID:20067610
4. http://www.ncbi.nlm.nih.gov/gene (NCBI gene search)
This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison