Model organisms:
Model organisms are important in order to research on a certain gene of interest. By using model organisms, researchers are able to create mutations in the gene being studied and then study the resulting phenotypes. The findings from these types of experiments can then be used to predict how the gene is functioning in humans and can grant us insight to the cause of a certain genetic diseases.
The ATM gene has been well conserved through evolutionary time as it is present in a wide variety of model organisms commonly used in genetic research. This has allowed researchers to do a great deal of research in order to explore the function of ATM and how it is linked to certain diseases. This page describes the phenotypes in three common model organisms, mouse, zebrafish, and C.elegans, which harbor ATM mutations. Visit the homology and phylogeny pages for more information regarding ATM and other organisms.
The ATM gene has been well conserved through evolutionary time as it is present in a wide variety of model organisms commonly used in genetic research. This has allowed researchers to do a great deal of research in order to explore the function of ATM and how it is linked to certain diseases. This page describes the phenotypes in three common model organisms, mouse, zebrafish, and C.elegans, which harbor ATM mutations. Visit the homology and phylogeny pages for more information regarding ATM and other organisms.
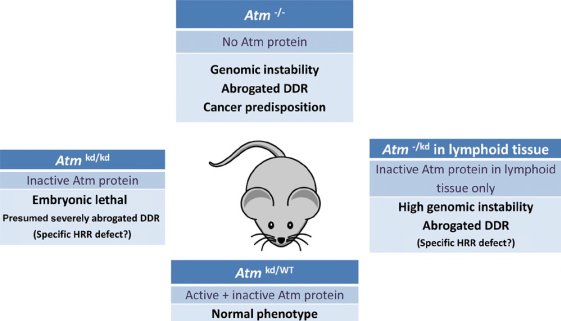
Mouse: In order to study phenotypic effects of ATM disruption, homologous recombination was used in order to create mice with ATM knocked out. Mice who were homozygous for the ATM mutation, labeled ATM(-/-), were found to be infertile due to complications in meiosis. Mice lacking ATM also showed to be predisposed to cancer and were also more sensitive to radiation. Furthermore, these mice also had problems with immune system development [1]. Researchers also created mice that harbored a mutation in the ATM gene that rendered its kinase activity ineffective, which was annotated as ATMkd (kd-kinase dead). These mice presented a more severe phenotype compared to ATM(-/-) animals, with the homozygous mutation being embryonic lethal. The reason for the more severe phenotype in ATM(kd/kd) mice compared to ATM(-/-) mice is not well understood but it is hypothesized that it could be due to the fact that kinase dead ATM has an inhibitory effect on homologous recombination, a process that is crucial during embryonic development [2].
In both mutants, V(D)J (variable, diversity, and joining) recombination, a process important in the development of lymphocytes that includes the development of double stranded breaks in DNA, was not as effective. This observation could be a possible explanation to the complications in immune development seen in ATM mutants. Figure 1 gives an overview of the different phenotypes in mice with ATM aberrations [3]. |
Figure 1: The different phenotypes seen in mice with the respective genotype. A (-) signifies the gene has been knockout while a kd signifies the presence of a kinase dead ATM protein. WT=wild type [3]
|
zeBRAfish:Figure 2: Zebrafish embryos injected with ATM morpholino in order to knockdown ATM. The embryos on the right are the controls that were not injected with morpholinos while the embryos on the left were. Note how the embryo injected with a higher MO concentration has a more severe phenotype. zATM MO= zebrafish ATM morpholino, Cont MO= control morpholino, 8 Gy= radiation does in Gray, pmol= concentration of MO [4].
|
In order to study the effects of ATM deficiency in zebrafish, morpholinos were used to inhibit the expression of ATM. It was found that zebrafish embryos that were injected with these morpholinos were more sensitive to radiation, which is consistent with other ATM studies in mice. These zebrafish showed gross developmental defects compared to wild type embryos exposed to radiation (figure 2) which resulted in lethality. It was also observed that higher concentrations of morpholino injections lead to more severe phenotypes and embryos that arrested earlier in development. These findings further support the importance of ATM in early development.
The zebrafish homolog of ATM was determined to have a high amount of identity with the functional domains of the human ATM protein. This conservation is important because it allows researchers to create mutants that, presumably, would result in a phenotype similar to human ATM mutations [4]. |
CaenorhaBditis elegans / RNAi experiments:
|
For C.elegans, it was found that worms with ATM defects showed phenotypes that were consistent with the presence of genomic instability after being exposed to radiation. These phenotypes include reduced viability after radiation exposure, Him phenotype, and sterility. Also, telomere maintenance seemed to be disrupted as well, due to the observation of increased chromosomal fusion events in ATM mutants [5].
When searching the RNAi database for RNAi phenotypes in C.elegans, it was found that the resulting phenotypes were wild type. Seeing as ATM has a role in nervous system development (ATM was shown to be involved in brain development, see Gene Ontology page), it is likely that is refractory to RNAi which could explain why the wild type result that was observed. In order to fix this problem, hairpin RNAi could be used, as the transgene method of RNAi delivery seems to be more effective against neuronal genes [6]. |
Figure 3: C.elegans that has been stained with a certain type of dye for imaging.
|
Discussion:
Knowing the resulting phenotypes from ATM mutations is important when trying to decide what model organism to use when studying ATM. Mouse seems to be the most desirable candidate as it is a mammal that contains a circulatory system, making it easy to model how ATM mutations affect leukemic cells. However, zebrafish is also a good organism to use as it is still a vertebrate and has a relatively large brood size and short generation time, as well as a primitive circulatory system. C.elegans is probably not the best model organism when looking for ATM’s connection to CLL as this organism does not have a circulatory system. Seeing as CLL involves the blood it is more beneficial to focus on organisms that have a circulatory system.
For this project, the proposed future directions uses mouse as the model organism because there have been a variety of ATM mutants already generated. Furthermore, conditional knockouts can be made in order to study the effect of ATM mutations in the circulatory system alone, without knocking out the gene in the rest of the animal.
For this project, the proposed future directions uses mouse as the model organism because there have been a variety of ATM mutants already generated. Furthermore, conditional knockouts can be made in order to study the effect of ATM mutations in the circulatory system alone, without knocking out the gene in the rest of the animal.
References:
1. Xu, Y., Ashley, T., Brianerd, E. E., Bronson, R. T., Meyn, M. S., & Baltimore, D. (1996). Targeted disruption of ATM leads to growth retardation, chromosomal fragmentation during meiosis, immune defects and thymic lymphoma. Genes and Development, 10, 2411-2422. doi:10.1101/gad.10.19.2411
2. Yamamoto, K., Wang, Y., Jiang, W., Liu, X., Dubois, R. L., Lin, C.-S., . . . Zha, S. (2012). Kinase-dead ATM protein causes genomic instability and early embryonic lethality in mice. Journal of Cell Biology, 198(3), 305-313. doi:10.1083/jcb.201204098
3. Shiloh, Y., & Ziv, Y. (2012). The ATM protein: The importance of being active. The journal of cell biology, 198(3), 273-275. doi:10.1083/jcb.201207063
4. Imamura, S., & Kishi, S. (2005). Molecular cloning and functional characterization of zebrafish ATM. The International Jounal of Biochemistry& Cell Biology, 37(5), 1105-1116. doi:10.1016/j.biocel.2004.10.015
5. Jones, M. R., Huang, J. C., Chua, S. Y., Baillie, D. L., & Rose, A. M. (2012). the atm-1 gene is required for genome stability in Caenorhabditis elegans. Molecular Genetics and Genomics, 287(4), 325-335. doi:10.1007/s00438-012-0681-0
6. Sugimoto, A. (2004). High-throughput RNAi in Caenorhabditis elegans: genome-wide screens and functional genomics. Differentiation, 72(2-3), 81-91. doi:10.1111/j.1432-0436.2004.07202004.x
2. Yamamoto, K., Wang, Y., Jiang, W., Liu, X., Dubois, R. L., Lin, C.-S., . . . Zha, S. (2012). Kinase-dead ATM protein causes genomic instability and early embryonic lethality in mice. Journal of Cell Biology, 198(3), 305-313. doi:10.1083/jcb.201204098
3. Shiloh, Y., & Ziv, Y. (2012). The ATM protein: The importance of being active. The journal of cell biology, 198(3), 273-275. doi:10.1083/jcb.201207063
4. Imamura, S., & Kishi, S. (2005). Molecular cloning and functional characterization of zebrafish ATM. The International Jounal of Biochemistry& Cell Biology, 37(5), 1105-1116. doi:10.1016/j.biocel.2004.10.015
5. Jones, M. R., Huang, J. C., Chua, S. Y., Baillie, D. L., & Rose, A. M. (2012). the atm-1 gene is required for genome stability in Caenorhabditis elegans. Molecular Genetics and Genomics, 287(4), 325-335. doi:10.1007/s00438-012-0681-0
6. Sugimoto, A. (2004). High-throughput RNAi in Caenorhabditis elegans: genome-wide screens and functional genomics. Differentiation, 72(2-3), 81-91. doi:10.1111/j.1432-0436.2004.07202004.x
This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison